Scribe Version 3.6
Scribe is the BrainMap application that is used to extract coordinates and meta-data from a published functional neuroimaging study and enter it in our database.

Database Content
The BrainMap project is committed to coding all fMRI and PET studies that publish their results in the form of Talairach or MNI coordinates. We have a team of staff and students in San Antonio that code papers for BrainMap on a daily basis. However, the body of literature is quite large. While we have made significant progress, there remains much to be done. If there are any papers that you wish to be included in BrainMap for your own analysis purposes, then we ask you to download Scribe and code those papers yourself.
The first 300 papers entered into BrainMap were chosen by taking 10 highly cited articles from 30 of the most prolific and respected functional neuroimaging laboratories in the world. Thereafter, papers were selected for BrainMap entry according to a series of meta-analyses performed at the Research Imaging Institute on various cognitive domains such as working memory, audition, executive function, pain, language, etc. Currently, database submissions are accepted either from students and staff at the RII or BrainMap users from the neuroimaging community.
Database Submission
The first panel of the Scribe interface is entitled "Citation", and the last is entitled "Feedback". The interface is designed to enter data from the left to the right.
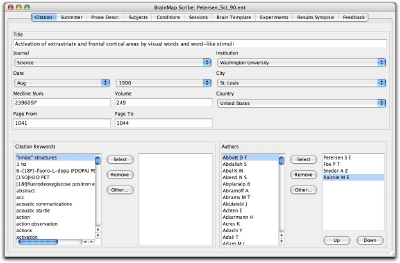
After all your meta-data and coordinates have been entered, save your submission by clicking on the top left program menu: File → Save As. Enter the file name in the following format: "Author_Journal_Year.ent". If you are on a PC, be sure and type in the file extension. Journal names should be abbreviated and years listed as the last two digits of the published year, for example, "Lee_HBM_02.ent".
When you are finished, email your BrainMap file and a .pdf of the original article to brainmap.org@gmail.com for review and insertion in the database.
Meta-Data Coding Scheme
BrainMap uses a standardized system of describing neuroimaging papers, termed the BrainMap Taxonomy. A citation section contains Authors, Journal Name and Year of Publication. A subject section has Diagnoses, Age, Handness and Native Language. Experiments are created by contrasting either functional conditions or VBM data. Hypothesis and experimental design are found in the Prose Description. The taxonomy page is a thorough reference point for understanding the coding scheme and essential for correctly coding papers.
In the case of functional studies, there are three experiment-level meta-data fields in our taxonomy that are extremely important to understanding the experiment and the isolated contrast:
Context
Context is defined as the purpose for which an experiment was performed, classified by the type of effect sought. Multiple contexts may apply for a given experiment. Possible choices here include: Normal Mapping, Age Effects, Disease Effects, Gender Effects, etc.
Paradigm Class
Paradigm Class is the experimental task isolated by your contrast. Typically, these paradigms have been used repeatedly by different researchers, with only minor changes made so that the essential structure is still recognizable. They have become widely known and accepted by brain imagers and generally have acquired informal (or formal) names. Multiple paradigm classes may apply for a given experiment. Possible choices here include: Action Observation, Delayed Match To Sample, Encoding, Episodic Recall, Oddball Discrimination, Stroop, Task Switching, etc.
Behavioral Domain
Behavioral Domain describes the categories and subcategories of mental operations likely to be isolated by the experimental contrast. Multiple behavioral domains may apply for a given experiment. Possible choices here include: Cognition.Memory.Working, Action.Execution.Speech, Cognition.Attention, Perception.Audition, etc.